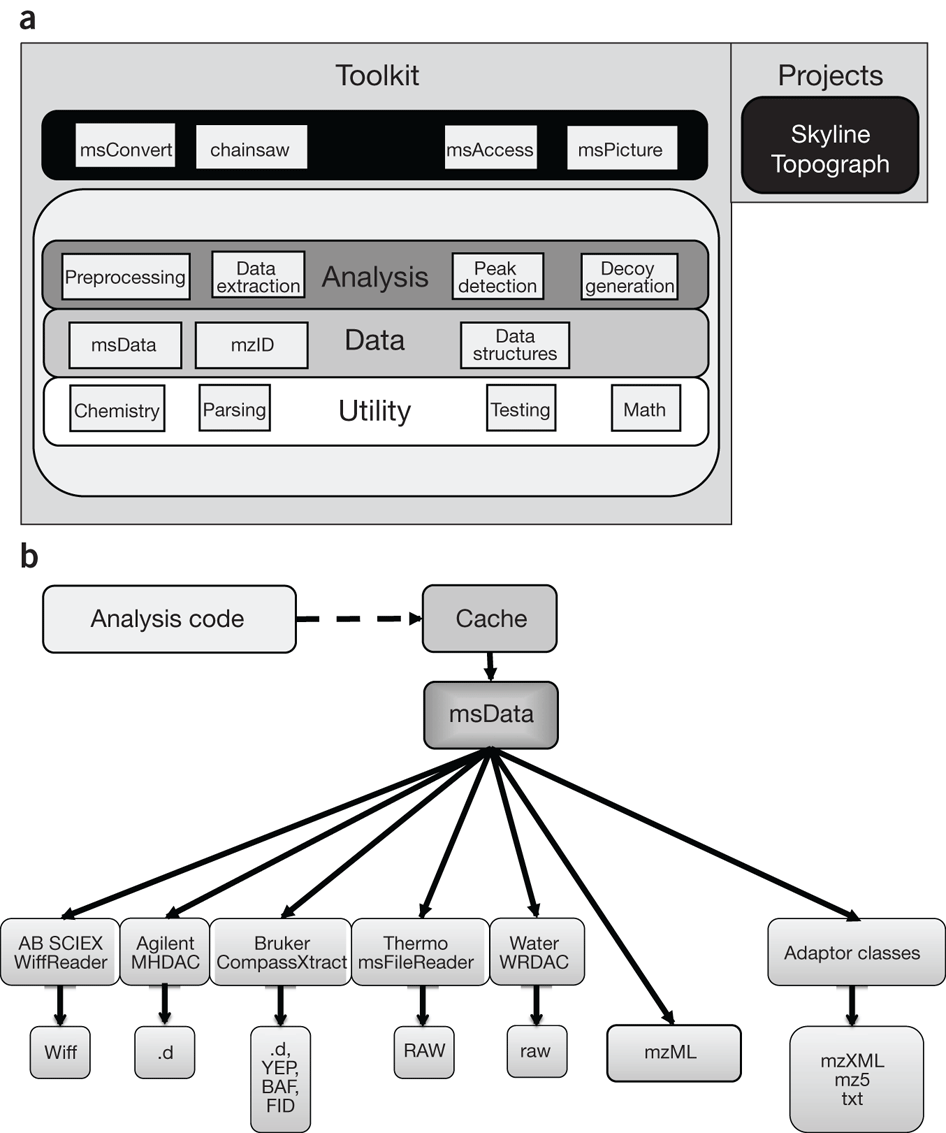
Methods to capture proteomic and metabolomic signatures from cerebrospinal fluid and serum of healthy individuals | Scientific Reports
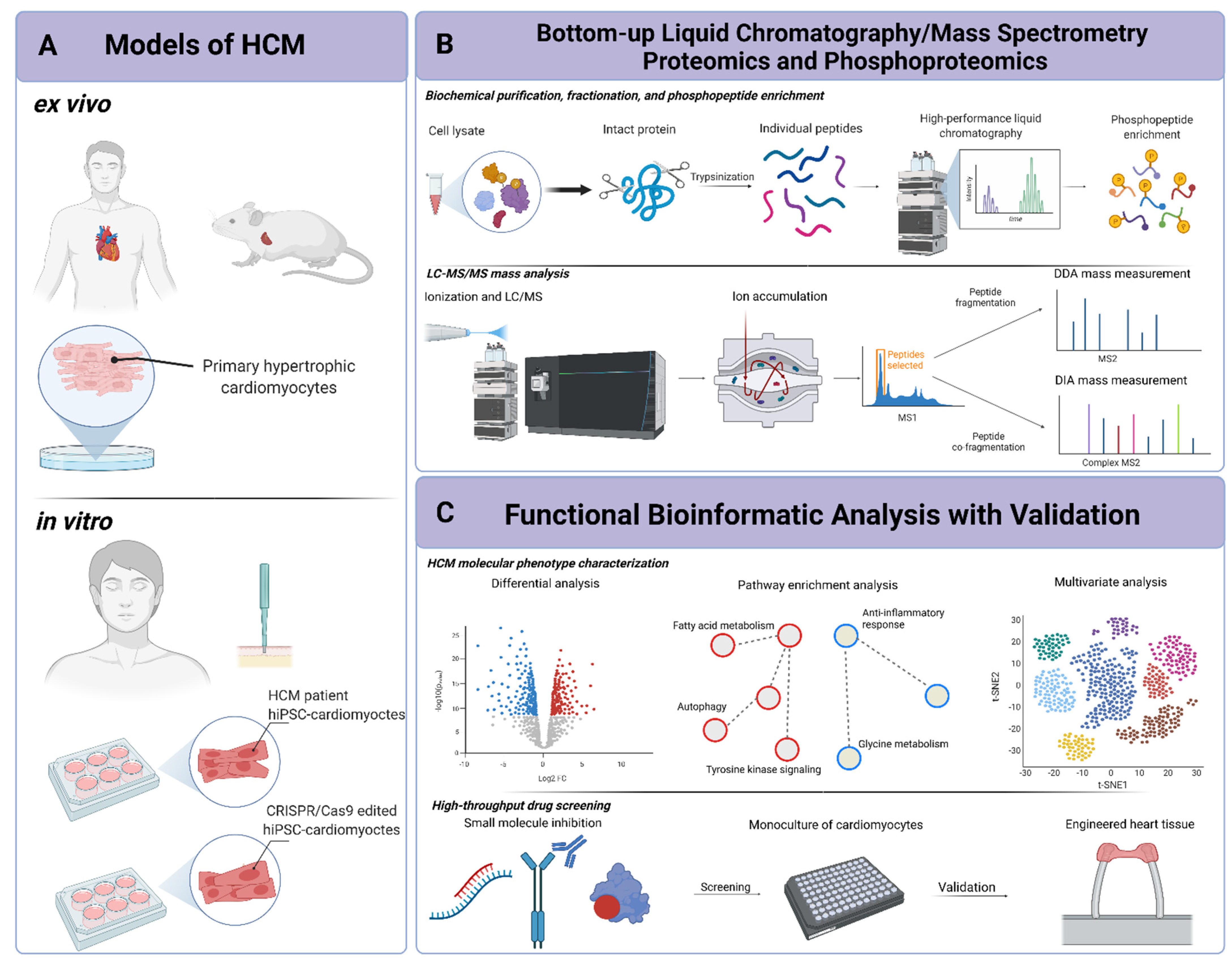
IJMS | Free Full-Text | Mass-Spectrometry-Based Functional Proteomic and Phosphoproteomic Technologies and Their Application for Analyzing Ex Vivo and In Vitro Models of Hypertrophic Cardiomyopathy
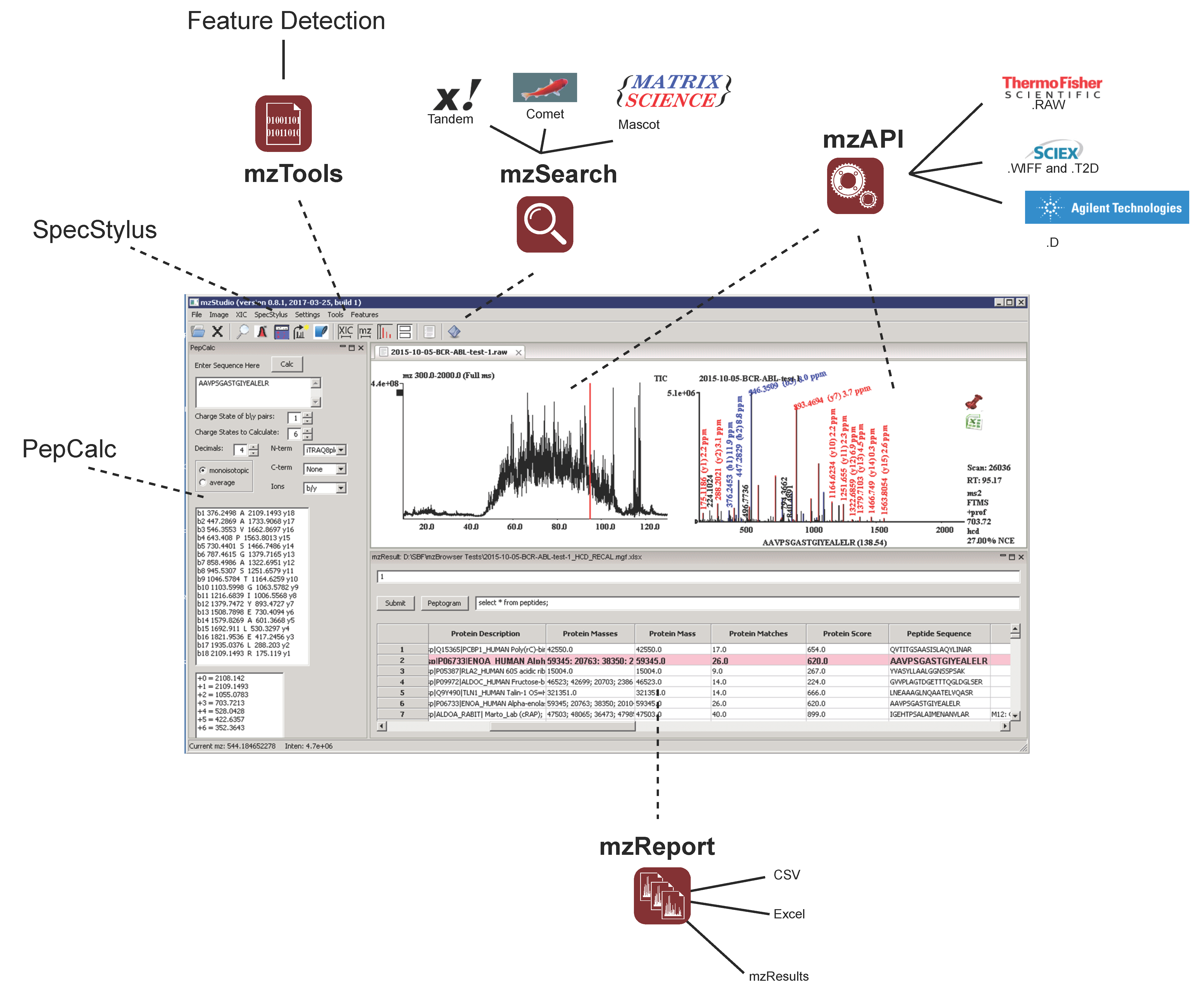
Proteomes | Free Full-Text | mzStudio: A Dynamic Digital Canvas for User-Driven Interrogation of Mass Spectrometry Data

Native Mass Spectrometry Imaging and In Situ Top-Down Identification of Intact Proteins Directly from Tissue | Journal of the American Society for Mass Spectrometry

Determining Alternative Protein Isoform Expression Using RNA Sequencing and Mass Spectrometry - ScienceDirect
![PDF] Single-platform 'multi-omic' profiling: unified mass spectrometry and computational workflows for integrative proteomics-metabolomics analysis. | Semantic Scholar PDF] Single-platform 'multi-omic' profiling: unified mass spectrometry and computational workflows for integrative proteomics-metabolomics analysis. | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/a99681b245131e0bc2523825fc0764e3de8c95c0/2-Table1-1.png)
PDF] Single-platform 'multi-omic' profiling: unified mass spectrometry and computational workflows for integrative proteomics-metabolomics analysis. | Semantic Scholar
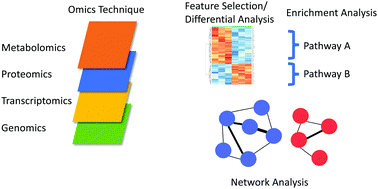
Single-platform 'multi-omic' profiling: unified mass spectrometry and computational workflows for integrative proteomics–metabolomics analysis - Molecular Omics (RSC Publishing)

Discovery of Protein Modifications Using Differential Tandem Mass Spectrometry Proteomics | Journal of Proteome Research